Overview
The team at the IGI Clinical Laboratory identified 100s of positive Sars-Cov-2 patient samples every week but didn’t have the bandwidth to build up the robust infrastructure to sequence these samples.
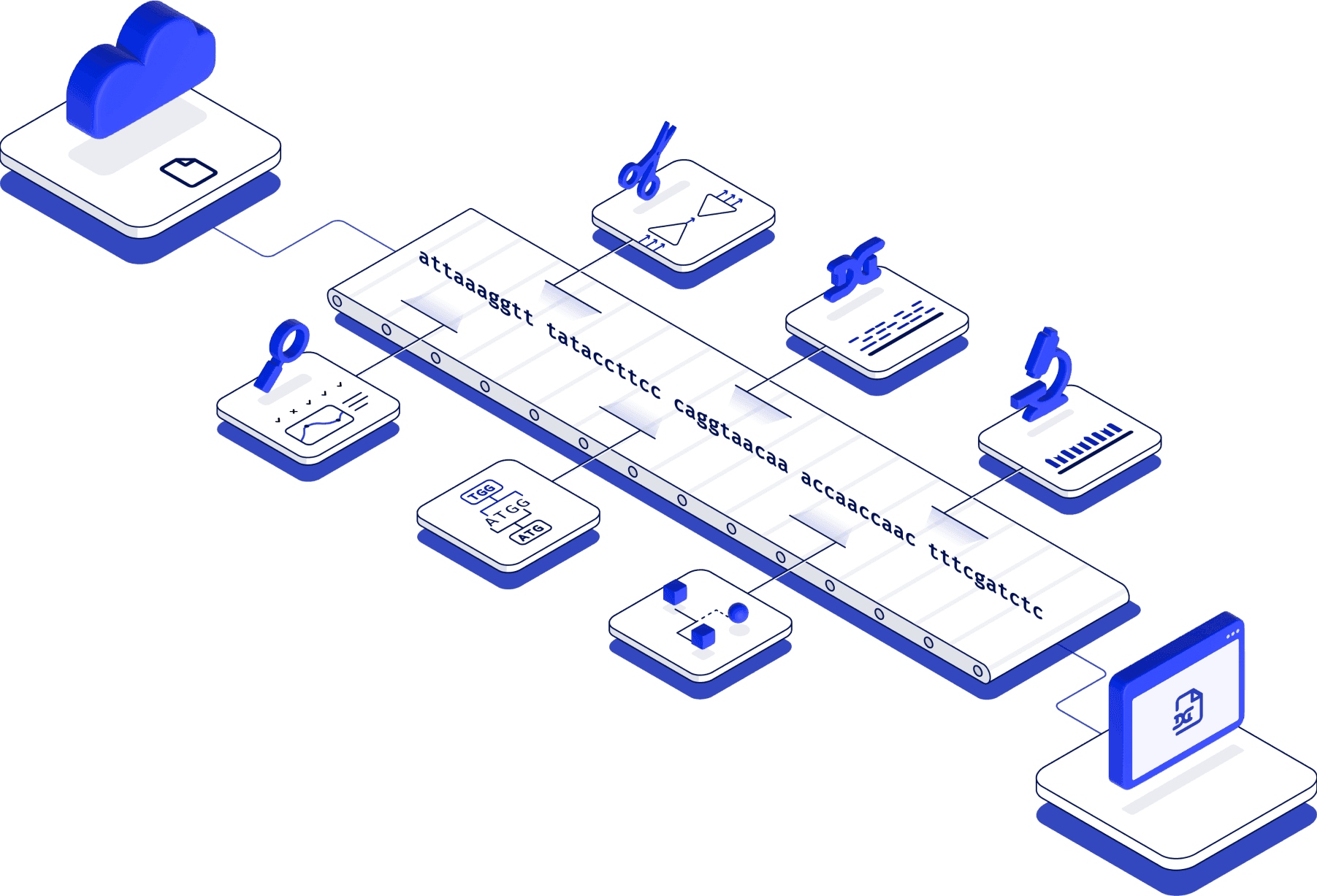
Latch teamed up to help them build a robust computational pipeline on the Latch platform which processes batches of several hundred samples in parallel. The result is a version-controlled and CLIA-compatible output that the testing personnel can use directly for patient care, or share with local public health departments for epidemiological use.
Through their collaboration with Latch the IGI CLIA:
- Automated processing of 10,000+ COVID samples through a UI for a single scientist to operate.
- The institute saved months of unneeded computing setup.
Challenges
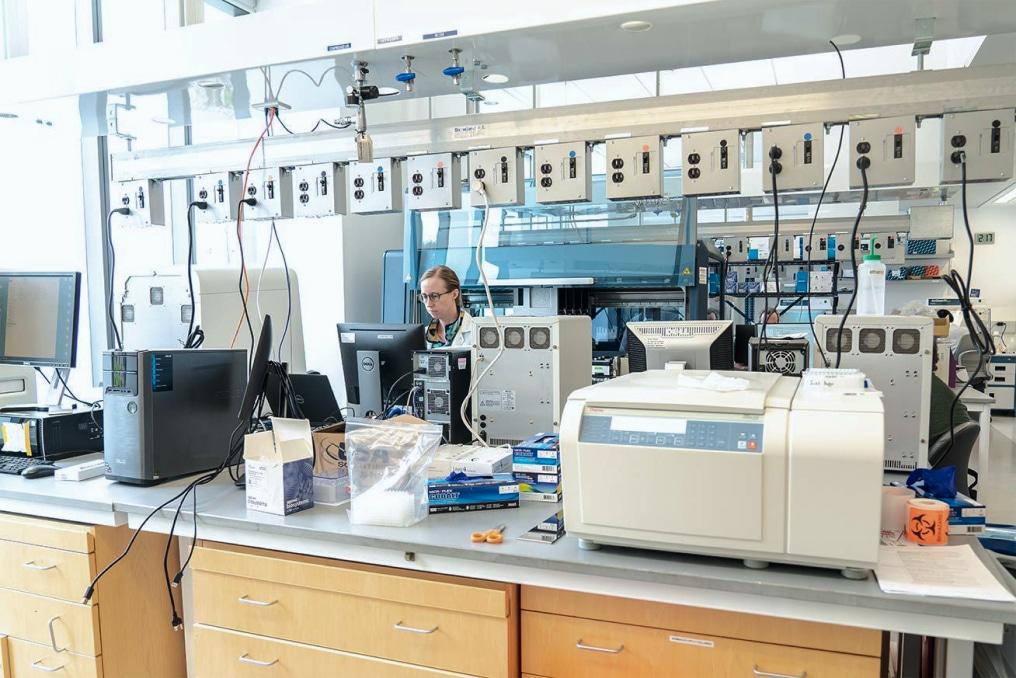
During the peak of the Sars-Cov-2 pandemic, the innovative genomics institute (the IGI) stepped into the scientific scene boldly to solve a bottleneck in diagnostic testing. Launching their first clinical lab in under ten weeks, the IGI planned to test thousands of patients across the Bay Area for different variants and clades of Covid-19.
However, the team at the research institute working within the CLIA (Clinical Laboratory Improvement Amendments) program was too busy at the bench to build complex infrastructure to process these samples.
Results
We worked with the great team at the CLIA IGI to build and host a robust and easily reproducible pipeline that processes 1000s of Covid-19 samples and produces patient-specific covid variants and clades.
CLIA scientists were able to simply import genomics sequencing data from Illumina BaseSpace, click “launch” and automatically run the whole Sars-Cov-2 pipeline with no bioinformatics needed.